What is Hardy-Weinberg equilibrium in AP Biology?
Hardy-Weinberg equilibrium is the no-evolution baseline in population genetics: if a population is not evolving at a locus, allele frequencies stay constant and genotype frequencies follow p + q = 1 and p² + 2pq + q² = 1. On the AP Biology exam you compare real counts to those predictions to spot violated assumptions.
In one sentence: Hardy-Weinberg equilibrium is the no-evolution baseline that predicts allele and genotype frequencies using p + q = 1 and p² + 2pq + q² = 1.
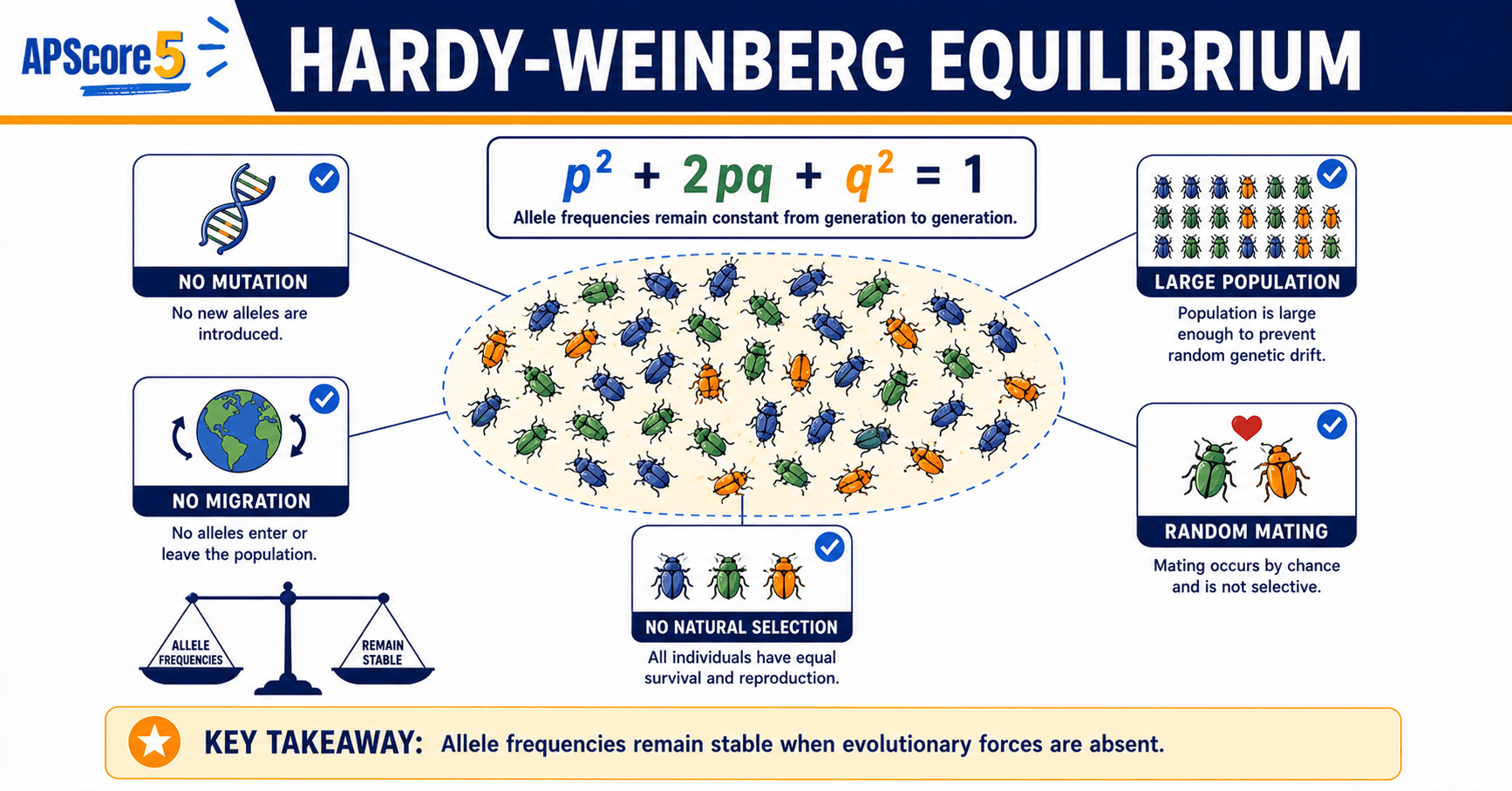
Plain-language snapshot
Hardy-Weinberg equation at a glance
Key idea: These two lines are the full Hardy-Weinberg equation pair tested as Hardy-Weinberg equilibrium AP Biology math—separate allele frequency (p, q) from genotype frequency (p², 2pq, q²).
Use p + q = 1 to connect the two allele frequencies in a two-allele population.
Allele frequencies sum to 100% of the gene pool.
Use when the population matches Hardy-Weinberg equilibrium assumptions.
Use p² + 2pq + q² = 1 to predict genotype frequencies when the population is in Hardy-Weinberg equilibrium.
AP trap: q is an allele frequency. q² is a genotype frequency.
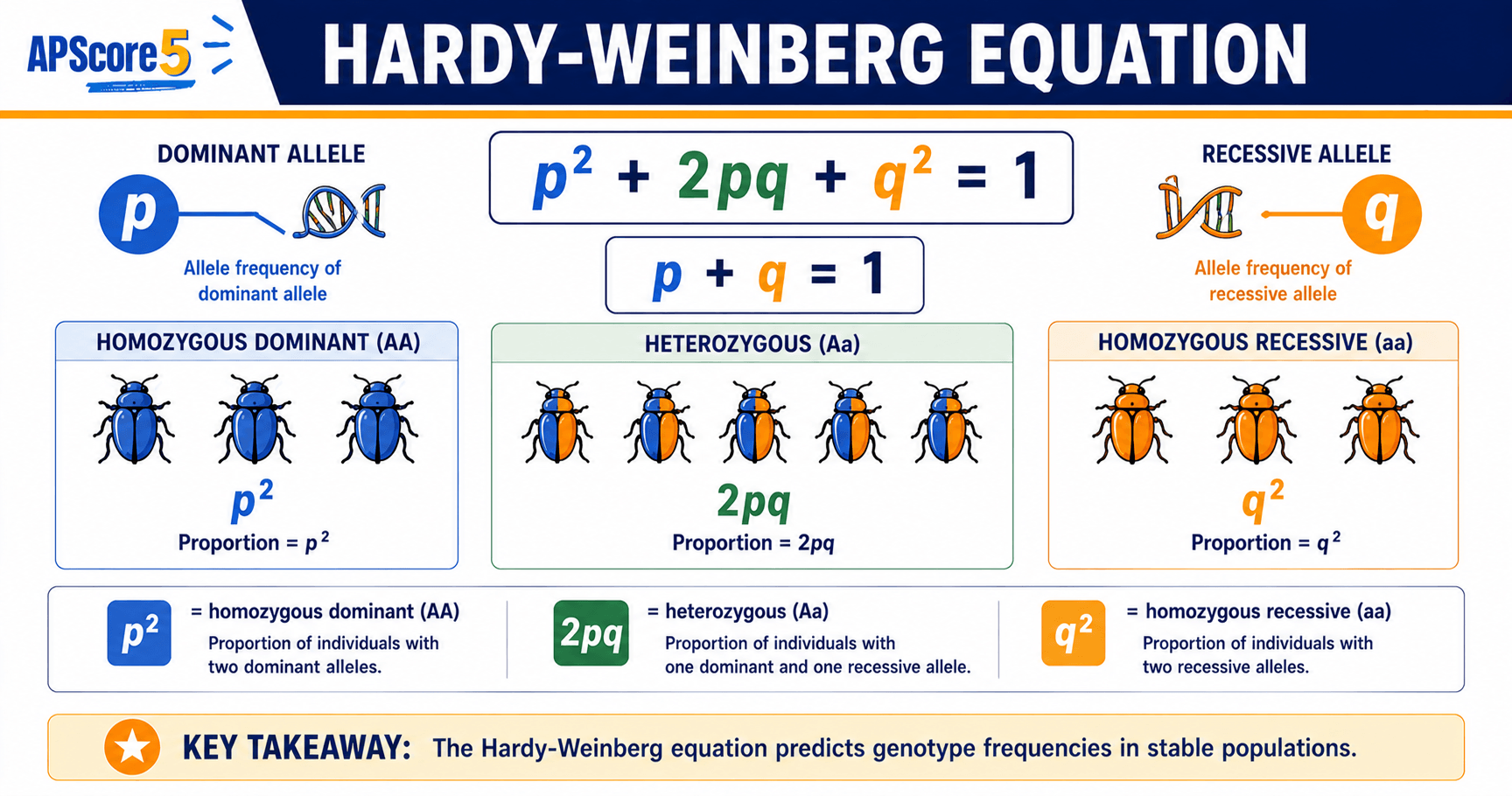
The two Hardy-Weinberg equations explained
Why it matters: Every Hardy-Weinberg practice problem boils down to these two lines—the Hardy-Weinberg equation pair on every formula sheet.
There are exactly two equations to memorize. Both stay manageable once you attach symbols to biology.
Equation 1: p + q = 1 (allele frequencies)
For a diploid autosomal gene with two alleles, every allele in the gene pool is either the dominant form or the recessive form. Frequencies must sum to 100%. If 70% of alleles are A, then p = 0.70 and q = 0.30 automatically.
Equation 2: p² + 2pq + q² = 1 (genotype frequencies)
Under random mating, multiply allele probabilities the same way you combine gametes in a Punnett square cross at population scale.
The three genotype bins are homozygous dominant (p²), heterozygous (2pq), and homozygous recessive (q²).
Most AP Biology Hardy-Weinberg problems start with q² because the recessive phenotype frequency equals q² when the trait is fully recessive.
| Term | Meaning | Example (p = 0.7, q = 0.3) |
|---|---|---|
| p² | Homozygous dominant frequency | 0.49 → 49% |
| 2pq | Heterozygous frequency | 0.42 → 42% |
| q² | Homozygous recessive frequency | 0.09 → 9% |
Why the 2 matters: heterozygotes arrive two ways—dominant allele from the maternal gamete or from the paternal gamete—so you must double pq.
Common mistake: Do not use 2pq as the recessive phenotype frequency—that slot is q² for aa.
Big picture: When mates combine alleles randomly and nothing else perturbs the gene pool, genotype frequencies match combinatorics from allele frequencies.
What are the five conditions for Hardy-Weinberg equilibrium?
Key idea: The five conditions for Hardy-Weinberg equilibrium are the checklist behind the Hardy-Weinberg null hypothesis—break one, and evolution or non-random mating can shift p and q.
Only when all five assumptions hold can you treat measured allele frequencies as stable and plug them into the equations without evolutionary correction.
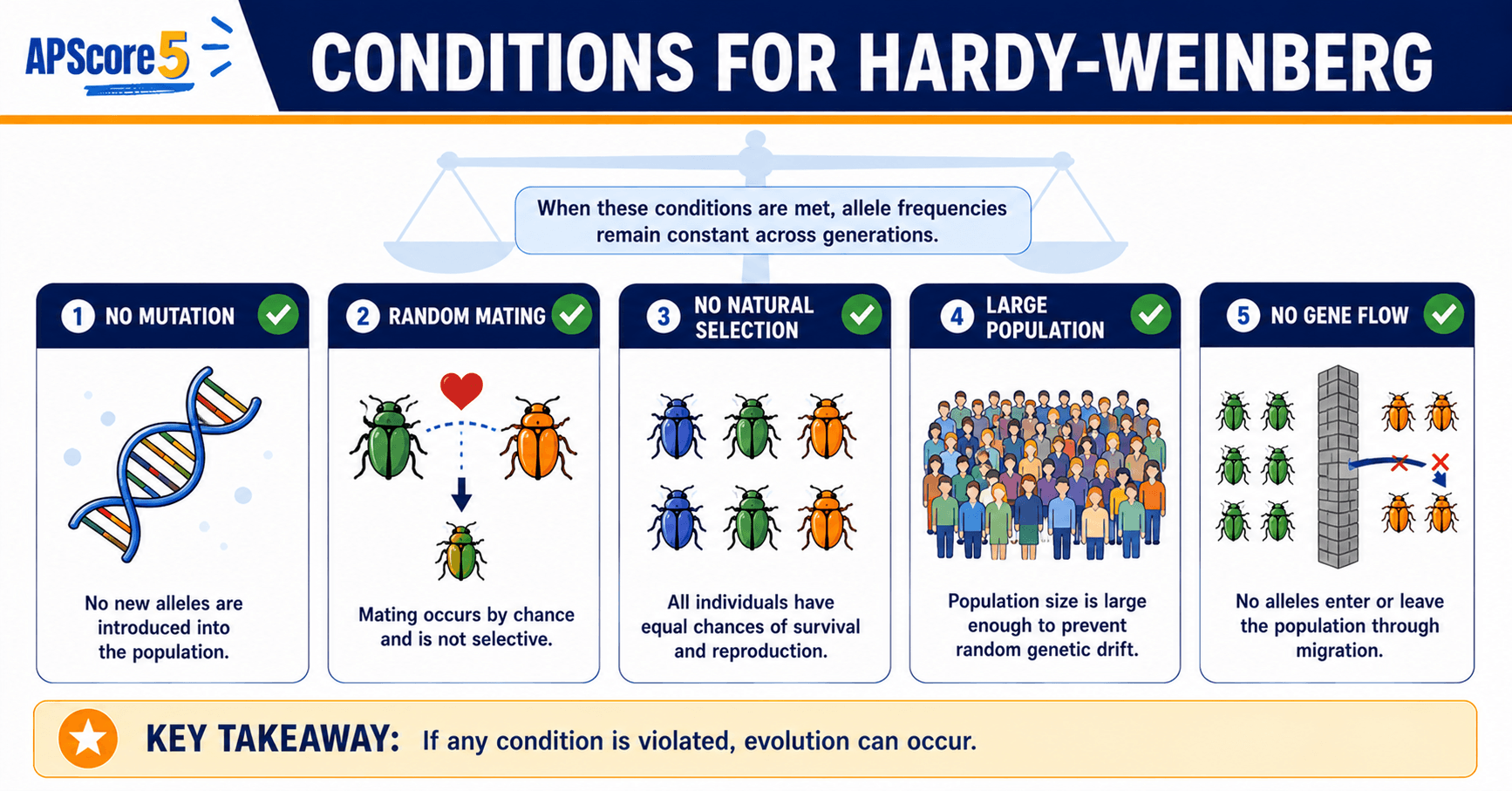
- No mutation: no new alleles enter the locus. Mutation rates are tiny per generation, but over long horizons mutation always occurs—another reason real populations only approximate HW.
- No migration: no gene flow in or out. Otherwise allele frequencies change because individuals carry different allele mixes.
- No natural selection: every genotype survives and reproduces equally. Selection breaks this instantly when some phenotypes leave more offspring.
- Very large population: genetic drift is negligible. Small populations fluctuate by chance even without selection.
- Random mating: individuals pair without respect to genotype at this gene—no inbreeding, no assortative mating, no sexual selection tied to the trait.
Memory hook: think “MMSDR” — Mutation, Migration, Selection, Drift (small population), Random mating. Break one → evolution can proceed.
AP trap: Hardy-Weinberg is a null model. It does not mean real populations never evolve—it means “Here is what we expect if evolution were absent at this locus.”
Concept link: Those five violations match the five major evolutionary mechanisms AP Biology expects you to connect—mutation, gene flow, selection, drift, and non-random mating.
How to solve Hardy-Weinberg problems step by step
Score booster: Say whether your final value is a frequency (0–1), a percent, or a count of individuals—AP readers reward explicit labels.
Most Hardy-Weinberg practice problems hide key information inside the recessive phenotype because aa individuals reveal q² directly—core how to solve Hardy-Weinberg problems logic.
- Find q² from the recessive phenotype.
- Take the square root to get q.
- Use p = 1 − q.
- Calculate p² and 2pq (carrier frequency when asked).
- Check that all frequencies add to 1.
- Identify q² from the fraction showing the recessive phenotype (often stated as “1 in 10,000”).
- Take √q² to solve for q.
- Compute p = 1 − q.
- Expand to p² and 2pq if the prompt asks for carriers or homozygous dominants.
- Verify that p² + 2pq + q² = 1 within rounding tolerance.
Why q² first? Dominant phenotypes lump AA with Aa carriers. Without sequencing families you usually cannot separate those classes from surveys alone, but recessive phenotypes isolate aa individuals.
Common mistake: Do not round until the final answer—early rounding skews p, q, and 2pq.
Calculator discipline: Store intermediate values and avoid rounding until the final line—AP grading tolerances tighten when students chop digits early.
Alternate entry points
Some stems give carrier frequency (2pq) instead of q². Divide by two before guessing p and q—you still enforce p + q = 1 after you identify which quadratic solution fits biology. Other prompts specify heterozygote counts in a census; convert counts to frequencies before plugging into expectations.
When q² is unknown
If only parental allele totals appear, compute p and q directly from allele counts, then square them for genotype predictions. Always label whether you counted alleles or individuals before you multiply.
Chi-square tie-ins
Hardy-Weinberg chi-square AP Biology prompts compare observed genotype counts to expected p², 2pq, and q² counts. Large χ² with biological context points to violated assumptions—not sloppy math.
Pedigree crossover: when a problem mixes family trees with population frequencies, translate pedigree conclusions into allele guesses only after you confirm inheritance mode matches autosomal recessive logic.
Exam pacing: sketch your five-step rail on scratch paper before touching the calculator so graders see clear structure even when arithmetic pressure rises mid-section.
Worked example: cystic fibrosis allele frequency
Cystic fibrosis is autosomal recessive. Roughly 1 in 2,500 people of European ancestry are affected. Treat that proportion as q² when the population approximates random mating for this gene.
Step 1: q² = 1/2500 = 0.0004.
Step 2: q = √0.0004 = 0.02.
Step 3: p = 1 − 0.02 = 0.98.
Step 4: p² ≈ 0.9604, 2pq ≈ 2(0.98)(0.02) = 0.0392, q² = 0.0004.
Step 5: Sum ≈ 1.000 after rounding.
Interpretation: even though only ~1 in 2,500 children show disease, about 1 in 25 people carry one allele—heterozygotes vastly outnumber affected homozygotes. Recessive alleles persist because they hide in carriers.
Score booster: When you report how to find carrier frequency Hardy-Weinberg style, label 2pq clearly as carriers—not as “recessive frequency.”
Real-world examples of Hardy-Weinberg in populations
Use this table to connect clinical prevalence (q²) to allele odds (q) and how to find carrier frequency Hardy-Weinberg estimates (~2pq).
| Disease / trait | q² (affected) | q (allele) | Carrier ~2pq |
|---|---|---|---|
| Cystic fibrosis (Europeans) | 1/2,500 | ~0.02 | ~1 in 25 |
| Phenylketonuria | 1/10,000 | ~0.01 | ~1 in 50 |
| Tay-Sachs (Ashkenazi) | 1/3,600 | ~0.0167 | ~1 in 30 |
| Sickle cell (parts of West Africa) | highlights selection | elevated q | heterozygote advantage |
Sickle cell caveat: malaria zones maintain the sickle allele partly because heterozygotes survive better—that is heterozygote advantage, so selection violates Hardy-Weinberg even though the equations still help you interpret measurements.
How Hardy-Weinberg appears on the AP Biology exam
Hardy-Weinberg questions reward careful algebra tied to biological meaning—expect both multiple-choice and FRQ-sized prompts.
AP Exam Answer Template
If observed genotype frequencies differ from Hardy-Weinberg predictions, at least one assumption is violated. That means mutation, migration, natural selection, genetic drift, or non-random mating may be changing allele frequencies.
In multiple-choice questions
- Given q², report q, p, 2pq, or p² without skipping steps.
- Identify which assumption fails in a scenario (migration event, bottleneck, mate choice).
- Distinguish allele equations from genotype equations.
In free-response questions
- Compare observed genotype counts to expected HW proportions using χ²-style reasoning when provided.
- Predict allele shifts when selection removes recessive phenotypes from breeding.
- Explain heterozygote advantage using malaria + sickle cell evidence.
Common stimulus types
Pedigrees paired with population surveys, allele frequency timelines, tables of observed versus expected genotypes, and word problems about carriers.
AP writing template
State given information → solve algebra → interpret in biological words → connect to an evolutionary mechanism or violated assumption.
Example: “q² = 0.01, so q = 0.1 and p = 0.9. Carriers equal 2pq = 0.18 even though only 1% show the recessive trait.” Then connect selection or drift if data drift away from those expectations.
Test yourself in 5 seconds
In a population at Hardy-Weinberg equilibrium, 16% of individuals are homozygous recessive (q²). What is the frequency of carriers (2pq)?
Hardy-Weinberg equilibrium vs evolving population
Hardy-Weinberg equilibrium AP Biology contrasts with evolving populations: constant p/q versus measurable change—your FRQ evidence lives in those deviations.
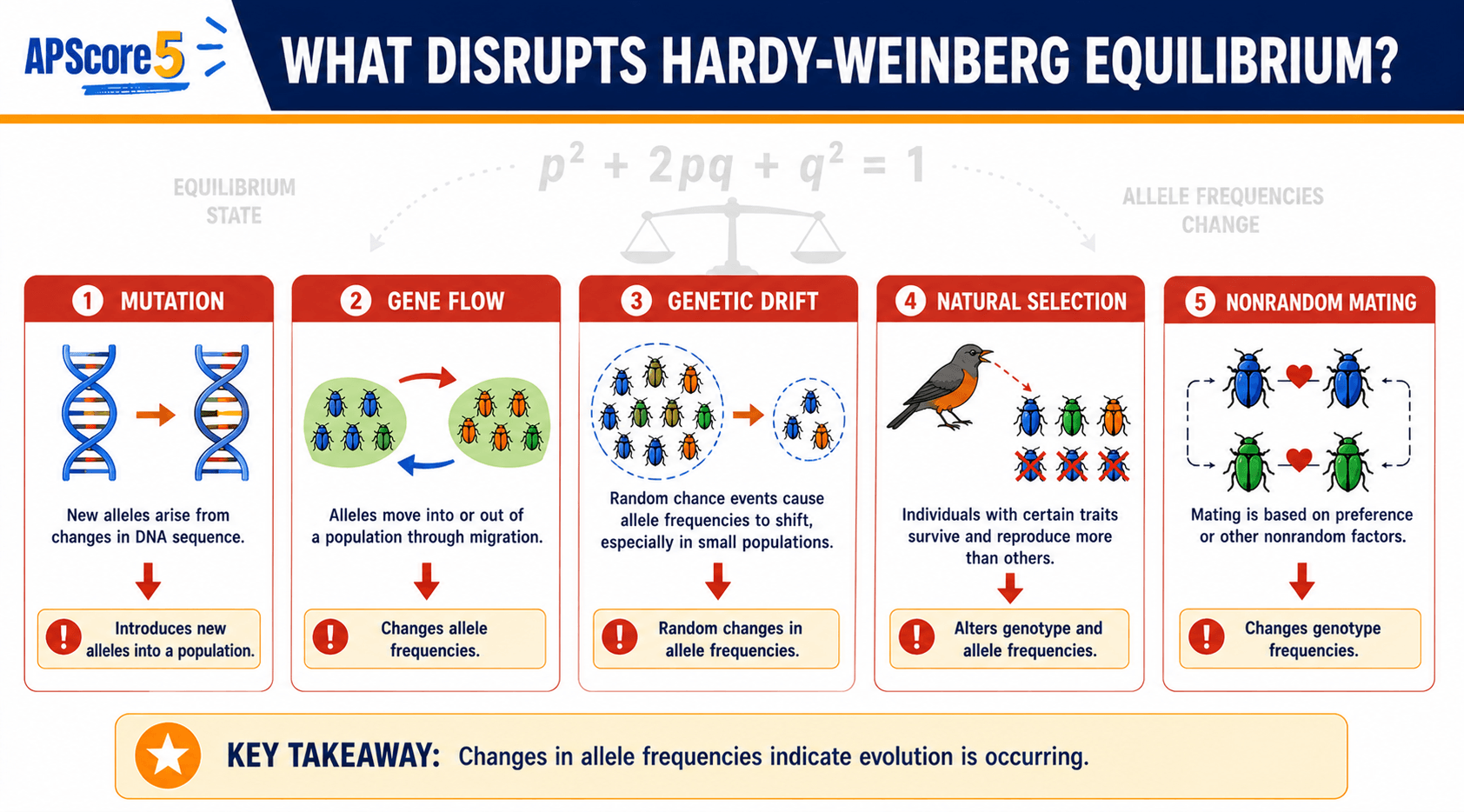
| Feature | HW equilibrium | Evolving population |
|---|---|---|
| Allele frequencies | Constant | Change across generations |
| Genotype fit | Matches p² + 2pq + q² | May diverge from expectation |
| Mutation | Negligible | Possible new alleles |
| Migration | None | Possible gene flow |
| Selection | Absent | May favor particular phenotypes |
| Population size | Very large | Any size—drift matters when small |
| Role on the exam | Null baseline | Typical real-world case |
Use deviations from HW expectations as evidence that evolutionary processes pull allele frequencies away from the static baseline.
Common AP Bio mistakes with Hardy-Weinberg
- AP trap: q is an allele frequency; q² is a genotype frequency—square roots bridge phenotype data to q.
- Mixing p with p²: alleles use p and q; genotypes square those values.
- Dropping the 2 in 2pq: heterozygotes require both parental orders.
- Starting from p² without justification: unless data separate AA from Aa, begin with q² when possible.
- Calling HW “more evolved”: equilibrium means no evolution at that locus.
- Assuming dominance changes algebra: dominance affects phenotype displays, not HW frequencies until selection acts.
- Counting heterozygotes inside q²: only aa individuals belong in q².
- Skipping the sum check: if frequencies fail to add to 1, hunt for arithmetic mistakes.
Why Hardy-Weinberg is the null hypothesis for evolution
Hardy-Weinberg null hypothesis framing: expect predicted genotype frequencies unless data reject the model.
Think of Hardy-Weinberg as the prediction you make when nothing evolutionary happens. Real studies compare observed genotype counts with expectations—large deviations spark investigations into selection, drift, migration, mutation, or mating structure.
This connects directly to AP Biology Unit 7 natural selection content on mechanisms that reshape allele pools over time.
For Mendelian vocabulary refresh, revisit AP Biology Unit 5 Heredity. For worked grid arithmetic before you square alleles, open Punnett square step-by-step practice—the same probability logic at individual-cross scale.
Hardy-Weinberg exam workshop
Translate words into symbols fast
Carrier prompts
Whenever a stem mentions carriers without naming genotypes, assume heterozygotes and map them to 2pq. Double-check whether the trait is recessive—dominant disorders invert which genotype you can count directly.
Frequency vs count
Convert population counts to decimals before squaring or square-rooting. If 400 individuals in 10,000 show the recessive phenotype, q² = 0.04—not 400.
Challenge drills
Selection overlays
If a stimulus removes aa individuals before reproduction, predict q decreases each generation even though HW equations describe the immediate parental pool before selection.
Sampling noise
Small samples deviate from expectations even without evolution. Exam writers sometimes hint at finite sample sizes—mention drift when populations are tiny islands or founder events appear.
Numeric hygiene
Guardrail checks
After solving, confirm p and q remain between 0 and 1 and genotype frequencies stay nonnegative. If carriers exceed 50% while q² is tiny, verify you multiplied p by q twice.
Units on FRQs
Label whether answers are frequencies or counts. When prompts demand expected counts, multiply decimals by census totals and round sensibly.
Timed loop: write “given → q² → q → p → target genotype → biology sentence.” Repeat until each step takes seconds, not minutes.
Hardy-Weinberg flashcards
Every fifth card advance triggers an ad placeholder with a three-second countdown before the next card appears.
Hardy-Weinberg practice problems & AP Biology MCQs
Hardy-Weinberg FRQ practice
Prompt: A population of 10,000 deer mice shows brown coat (dominant B) and tan coat (recessive b). Researchers count 400 tan mice and assume Hardy-Weinberg equilibrium.
- (A) Calculate q and p. Show work.
- (B) Calculate expected counts of BB and Bb mice.
- (C) Twenty years later only 100 tan mice appear although census stays 10,000. Explain whether HW holds and name one mechanism.
- (D) Predict directional effects if hawks preferentially capture tan mice.
Sample 4-point response
(A) Tan mice are bb, so q² = 400/10,000 = 0.04. Thus q = 0.2 and p = 0.8.
(B) Expected BB = p²N = 0.64 × 10,000 = 6,400. Expected Bb = 2pqN = 0.32 × 10,000 = 3,200. Check: 6,400 + 3,200 + 400 = 10,000.
(C) HW is violated because q² dropped from 0.04 to 0.01, changing q from 0.2 to 0.1. Natural selection against tan mice or non-random mating could explain the shift.
(D) Predation removes recessive phenotypes from breeding pools, lowering q over time. Heterozygotes still hide b alleles, so recessive alleles linger.
Rubric: A — correct q and p with shown work. B — accurate genotype counts. C — identifies violation plus plausible mechanism. D — directional selection explanation referencing heterozygotes.
Hardy-Weinberg equilibrium FAQ
What is Hardy-Weinberg equilibrium?
It is the no-evolution baseline for one gene: allele frequencies stay constant and genotype frequencies follow p + q = 1 and p² + 2pq + q² = 1 when the five assumptions hold.
What are the two Hardy-Weinberg equations?
p + q = 1 sums allele frequencies for two alleles. p² + 2pq + q² = 1 sums expected genotype frequencies (homozygous dominant, heterozygous, homozygous recessive) under random mating.
What are the five conditions for Hardy-Weinberg equilibrium?
No mutation, no migration (gene flow), no natural selection, very large population size (negligible genetic drift), and random mating.
How do you solve Hardy-Weinberg problems?
Often estimate q² from the recessive phenotype, take the square root for q, use p = 1 − q, compute p² and 2pq, then confirm p² + 2pq + q² = 1.
How do you find p and q in Hardy-Weinberg?
If q² is known from aa frequency, q = √(q²) and p = 1 − q. If raw allele counts are given, divide each allele count by the total number of alleles in the sample.
How do you find carrier frequency in Hardy-Weinberg?
For autosomal recessive traits, carriers are heterozygotes—use 2pq after you know p and q.
Why do many Hardy-Weinberg problems start with q²?
Homozygous recessives usually show the recessive phenotype, so surveys isolate q²; dominant phenotypes mix AA and Aa unless you have extra data.
What does it mean if a population is not in Hardy-Weinberg equilibrium?
Observed genotype frequencies differ from p², 2pq, and q² expectations—mutation, migration, selection, drift, or non-random mating may be changing allele frequencies.
Is Hardy-Weinberg a null hypothesis?
Yes—it predicts genotype proportions if evolution is absent at that locus; departures mean at least one assumption fails.
How is Hardy-Weinberg tested with chi-square in AP Biology?
Multiply expected frequencies by sample size to get expected counts for each genotype class, run a chi-square goodness-of-fit test against observed counts, then interpret significant deviation with biology (not just “bad math”).
Why is there a 2 in 2pq?
Heterozygotes arise two ways (A from one parent and a from the other, or the reverse), so the heterozygote term doubles pq.
Is any real population actually at Hardy-Weinberg equilibrium?
Rarely perfect—real populations usually violate at least one assumption slightly; HW stays useful as a benchmark.
What is the connection between Hardy-Weinberg and natural selection?
Hardy-Weinberg assumes no selection. When survival or reproduction differs by genotype, observed frequencies drift away from p² + 2pq + q² predictions.
Why are recessive disease alleles common in carriers but rare in affected people?
Recessive alleles hide in heterozygotes—when q² is small, 2pq can still be much larger than q².
How does Hardy-Weinberg relate to Punnett squares?
Both model random union of gametes; a Punnett square is one cross, while Hardy-Weinberg applies the same probability rules across a whole randomly mating population.